Force of Infection is a reader-supported publication. To support our work and access our weekly, detailed epidemiology reports, consider upgrading.
This is the second installation in a series about respiratory viruses and their complexities. The first essay explained less common viruses like human metapneumovirus and adenovirus. Today, we're taking a deeper look at disease surveillance.
Why don’t I know what I have?
If you go to the doctor or hospital with what feels like a “cold,” there’s a set of decisions happening behind the scenes about whether to test you, and for what.
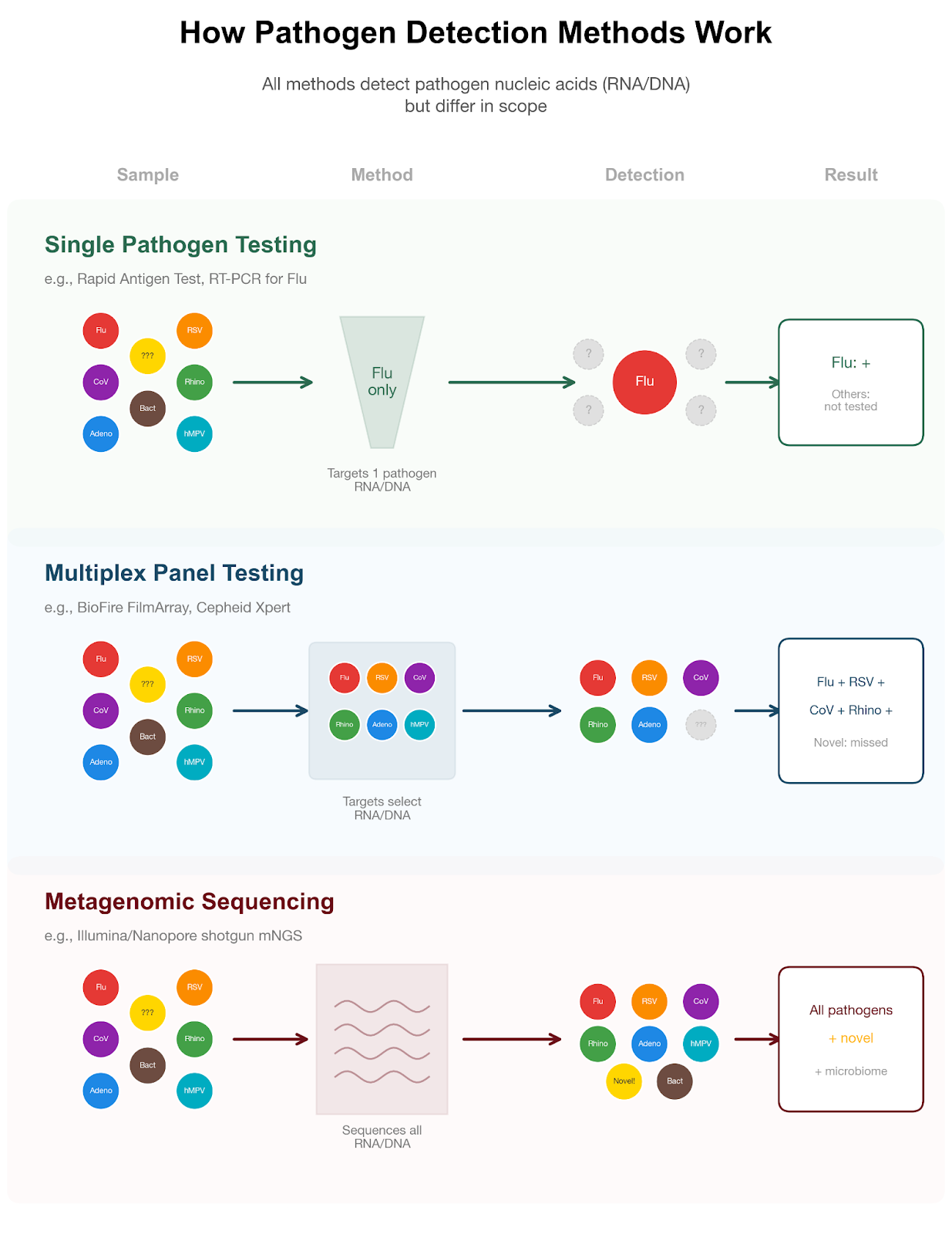
In most outpatient settings, testing is usually “targeted.” This means you might get a single test (for example, just for influenza), or a small “triplex” panel that checks for the most common viruses: SARS-CoV-2, flu, and RSV. These are the pathogens where a diagnosis can directly affect care or public health decisions.
Testing for a broader array of pathogens does exist, but is not routinely used. Some specialized multiplex (or, multi-pathogen) panels, like the BioFire FilmArray, can detect 20-25 pathogens at once. These panels cover not just influenza, RSV, but also the common human coronaviruses, rhinovirus/enterovirus, and the other uncommon pathogens described in part I of this series. However, these panels only detect known viruses included on the test, they cannot identify entirely new or emerging pathogens, which require genomic sequencing approaches. Many viruses, such as human metapneumovirus (hMPV) and parainfluenza (HPIVs), don’t have widely used standalone clinical tests and so they are typically only identified when these multiplex panels are run. Even then, results are sometimes reported at the group level, meaning the test detects a category of virus (for example, “parainfluenza virus”) without specifying the exact subtype (such as HPIV-1 vs. HPIV-3).
Broad panels are usually reserved for hospitalized patients or people at higher risk of severe disease. In those settings, identifying the pathogen can directly change care. For example, a confirmed influenza diagnosis may lead to antiviral treatment, while identifying a viral cause can support stopping unnecessary antibiotics. Results also guide infection control, such as isolating patients or grouping them with others who have the same infection, to prevent spread within the hospital.
For people who are otherwise healthy, however, knowing the exact virus rarely changes management. Most respiratory infections are treated with supportive care; rest, hydration, and symptom relief, regardless of the cause.
There are also practical constraints. Broad panels are more expensive than targeted tests, take longer to process, and may not be available in all care settings. As a result, testing is generally focused on situations where the results will meaningfully change what happens next.
The result is a gap between individual experience and medical knowledge. Many people get respiratory infections, but relatively few ever receive a definitive diagnosis.
How do we know what’s circulating?
Epidemiologists don’t rely on individual diagnoses to track respiratory viruses. Instead, they use a patchwork of surveillance systems, each capturing a different slice of what’s happening in the population.
- Lab-confirmed tests: Certain hospitals and diagnostic labs, called sentinel labs, regularly report results to public health agencies. These labs contribute data consistently, giving a steady sense of which viruses are circulating and how activity is changing over time.
- Syndromic surveillance: Some systems track symptoms rather than confirmed infections, such as emergency room visits for fever, cough, or influenza-like illness. These data systems are timely and cover more people, but less precise, since different viruses can cause similar symptoms. As long as care-seeking behavior is stable (i.e., people are visiting the emergency department at consistent rates, in contrast to times like a natural disaster when ED use may be impacted), these trends are highly informative.
- Multiplex testing data: Increasingly, a subset of patients are tested for many pathogens at once. These data fill in gaps between syndromic signals and lab-confirmed diagnoses, showing which specific viruses are contributing to observed trends.
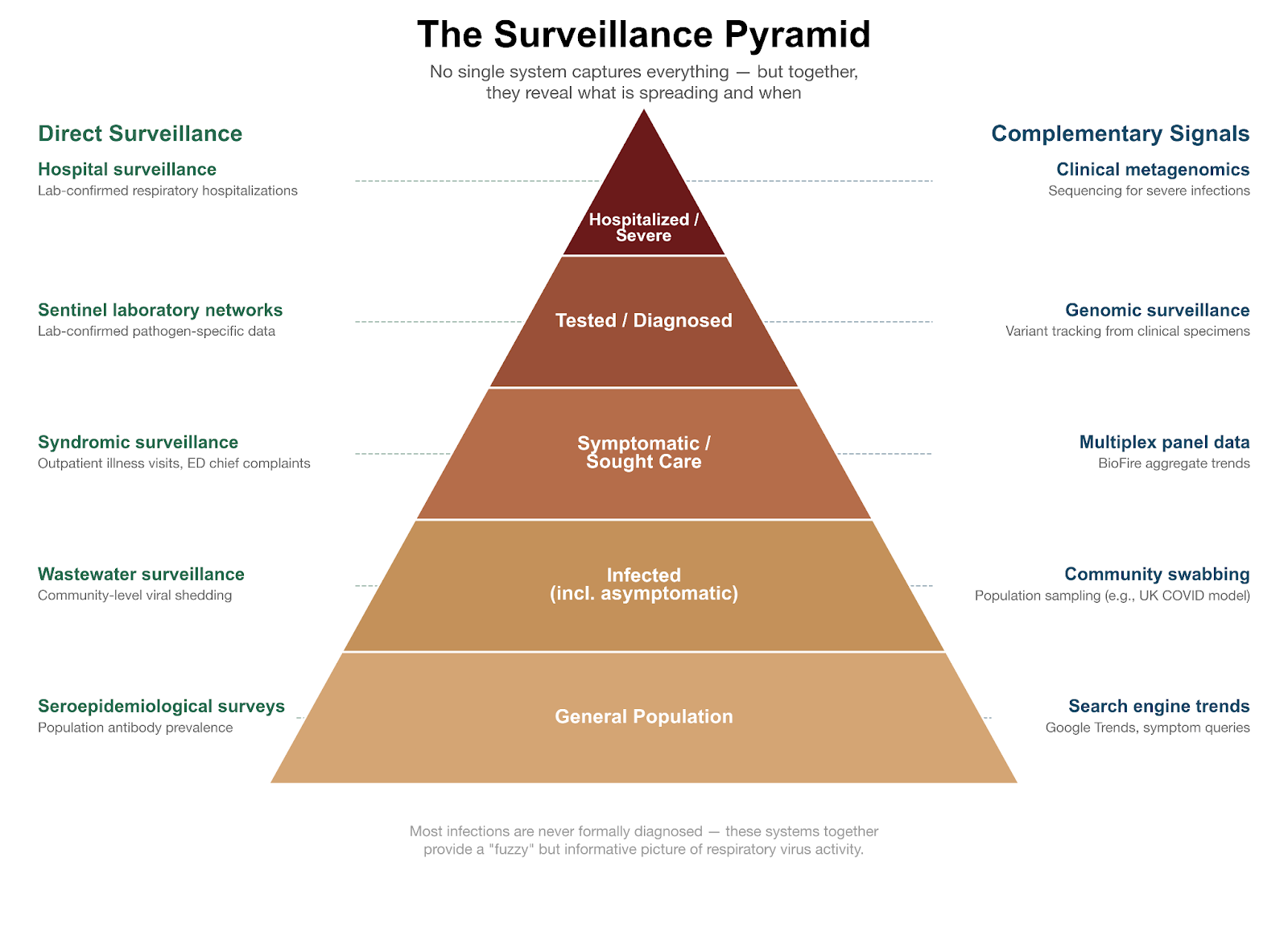
You can see how these systems work together: a rise in emergency visits for influenza-like illness may be the first signal of increasing respiratory activity. Sentinel labs then report which viruses are present, and multiplex panels can reveal whether RSV, hMPV, or other viruses are also contributing.
The result is a “fuzzy” but informative picture of virus activity. No single system captures everything, but together they allow us to track when respiratory illness is increasing or decreasing, how seasonal peaks compare to previous years, and which pathogens are likely driving current trends.
That said, not all viruses are tracked equally well. Surveillance is most robust for well-studied pathogens like influenza and RSV, where large, coordinated networks provide near real-time, geographically detailed data. For viruses like human metapneumovirus or parainfluenza, coverage is more limited, reporting is often delayed, and estimates are less precise.
There are also important limitations. Surveillance data are typically based on a subset of locations and may not reflect local conditions with high geographic specificity. Changes in testing practices or care-seeking behavior can influence what gets measured. And because most infections are never formally diagnosed, all of these systems observe only a fraction of total cases.
Even with these gaps, the overall trends are surprisingly reliable. While we may not know exactly what any one person has, we can still understand, at a population level, what is spreading and when.
The ideal vision for tracking respiratory viruses
One way to think about the limits of current systems is to imagine an ideal one. In a perfect scenario, a random fraction of the population would regularly submit nasal swabs for testing. Combined with broad molecular panels and genomic sequencing, this could provide a near real-time view of circulating viruses, including common, rare, and potentially novel pathogens.
During the Covid-19 pandemic, the UK came close to this ideal, but only for a single virus: SARS-CoV-2. Large-scale community swabbing programs randomly sampled people across the population and sequenced positive samples, giving an unprecedented view of how Covid-19 was spreading, including new variants. While highly informative, applying this approach to all respiratory viruses would be far more expensive and logistically complex.
Tools approaching the ideal
- Wastewater surveillance: Testing sewage for viral genetic material shed in stool, urine, or respiratory secretions provides near real-time, population-level trends. While most attention has focused on SARS-CoV-2, this infrastructure is increasingly used for influenza, RSV, adenoviruses, and other viruses. However, wastewater cannot identify individual cases, severity, or exactly who is affected, and detection can vary depending on population size and viral shedding.
- Air sampling: Collecting and analyzing airborne particles in indoor or high-risk environments (e.g., hospitals, schools, transportation hubs) allows for direct detection of viral genetic material suspended in aerosols. Air sampling can provide early warning of circulating respiratory viruses, including SARS-CoV-2, influenza, and RSV, especially in settings where transmission risk is high. Unlike wastewater, it is more spatially resolved and can reflect current airborne exposure risk. However, sensitivity depends heavily on sampling duration, air flow, and viral concentration, and detection does not necessarily indicate infectious virus or identify specific individuals.
- Digital data streams: Google Trends or other search activity can provide early signals of rising infections. Increases in symptom-related searches often precede lab-confirmed cases, offering a useful complement to traditional surveillance. These tools are early-warning signals, not precise diagnostics.
- Clinical metagenomics: The most comprehensive approach sequences all genetic material in a sample, detecting known and novel pathogens simultaneously. Unlike traditional targeted tests, it generates full viral genomes, which can be compared to other sequences to trace how a virus spreads, showing how it entered a community, how long it has been circulating, and, in studies with dense sampling, potentially even who infected whom. However, metagenomics is not widely used clinically. It is expensive, because each sample is processed individually with specialized sequencing machines and costly reagents. It is time-intensive, taking hours to days to generate sufficient data. And it is technically challenging, requiring advanced bioinformatics and expert interpretation, which makes it primarily a research tool or an option for unusual or severe infections rather than routine care.
Looking ahead
Longer term, our aspiration at Force of Infection is to see a more comprehensive and responsive surveillance system, one that captures not just the viruses epidemiologists already prioritize, but the broader ecosystem of respiratory pathogens that shape everyday illness. By combining digital signals, wastewater monitoring, and clinical metagenomics with the surveillance programs we already rely on, we could track respiratory viruses more quickly and accurately, filling in gaps and providing a much clearer picture of what’s circulating in communities. This integrated approach could make virus tracking faster, more informative, and more actionable for public health decision-making.
In the meantime, the next time you’re sick and test negative for the usual suspects, the most likely explanation is that you’ve encountered one of the many viruses we don’t routinely test for.
Force of Infection is a reader-supported publication. To support our work and access our weekly, detailed epidemiology reports, consider upgrading.
Gage Moreno, PhD, is a genomic epidemiologist who uses large-scale sequencing data to study respiratory virus transmission, focusing on how pathogens spread through communities and how we detect them.


